Day 5 — Reproducible Pipelines: Snakemake and Nextflow
Day 5: Reproducible Pipelines — Snakemake and Nextflow
This is Day 5 of a 6-part series: From Laptop to HPC: Scaling Computational Biology Workflows and 🧬 Day 62 of Daily Bioinformatics from Jojy’s Desk. In Day 4 we scaled to 200 samples in parallel with job arrays. Today: the final evolution — workflow managers that make the whole thing reproducible.
The Hidden Problem With Shell Scripts
You’ve spent four days building a powerful HPC toolkit. You can run single jobs, handle 200 samples in parallel, load modules, manage environments. Your shell scripts work.
Then this happens:
Six months later. A reviewer asks you to rerun the analysis with a slight parameter change. You open your scripts. You don’t remember which version of samtools you used. Your conda environment was updated since then. Two intermediate files are missing because a job silently failed and you didn’t notice. You rerun everything from scratch, but the results are slightly different. You don’t know why.
Or this:
A collaborator wants to run your pipeline on their cluster. You send them your SLURM scripts. They spend three days troubleshooting different module names, missing paths, and job submission order before giving up.
Shell scripts are not pipelines. They’re instructions. The difference matters.
A workflow manager solves this by:
- Tracking which outputs depend on which inputs (the dependency graph)
- Only re-running steps whose inputs have changed (incremental re-runs)
- Automatically restarting from the point of failure
- Explicitly documenting every tool version and parameter
- Running identically on laptop, HPC, and cloud — with one config change
What You’ll Learn Today
- Why dependency tracking matters and what a DAG is
- How to write a Snakemake pipeline from scratch
- How to write a Nextflow pipeline from scratch
- How to run both on HPC with SLURM integration
- Which tool to choose for your project
- A GitHub repository structure that makes your pipelines shareable
- Publishing checklist for reproducible bioinformatics
Part 1: The Dependency Graph (DAG)
Before writing code, let’s understand what workflow managers actually do.
In a bioinformatics pipeline, outputs of one step become inputs of the next:
raw_reads.fastq.gz
│
▼
[FastQC] → quality_report.html
│
▼
[Trimmomatic] → trimmed_reads.fastq.gz
│
▼
[Bowtie2] → aligned.sam
│
▼
[samtools sort] → aligned.bam
│
▼
[samtools index] → aligned.bam.bai
│
▼
[featureCounts] → counts.txt
This is a Directed Acyclic Graph (DAG) — a chain of steps where each depends on the previous.
With shell scripts, you manage this manually. With workflow managers, you declare what inputs produce what outputs, and the tool figures out the order automatically. More importantly, if aligned.bam already exists and trimmed_reads.fastq.gz hasn’t changed, the workflow skips the alignment step entirely.
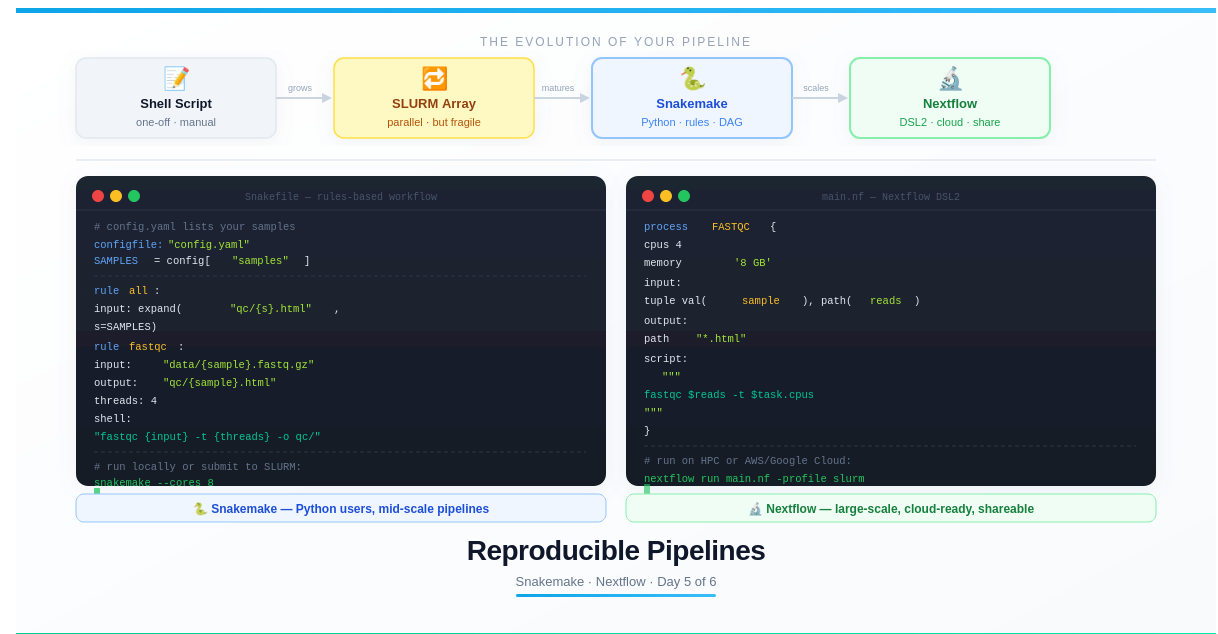
Part 2: Snakemake — Python-Native Workflow Manager
Snakemake is the most widely used workflow manager in academic bioinformatics. If you know Python, the syntax will feel natural immediately.
Core Concept: Rules
In Snakemake, you write rules. Each rule says: “given these inputs, run this command to produce these outputs.”
# The most basic possible Snakemake rule
rule fastqc:
input: "data/{sample}.fastq.gz"
output: "qc/{sample}_fastqc.html"
shell: "fastqc {input} -o qc/"
That’s it. Snakemake figures out:
- Which samples need processing (by matching the wildcard
{sample}) - Whether the output already exists (skip if yes)
- What order to run rules in (based on dependency chains)
A Complete RNA-seq Snakemake Pipeline
# Snakefile
# ─────────────────────────────────────────────
# Complete RNA-seq pipeline: FastQC → Trim → STAR → featureCounts
# ─────────────────────────────────────────────
configfile: "config.yaml"
SAMPLES = config["samples"]
REF = config["genome"]
GTF = config["gtf"]
# ── Rule all: defines the final outputs ──────
rule all:
input:
expand("qc/{s}_fastqc.html", sample=SAMPLES),
expand("bam/{s}.sorted.bam", sample=SAMPLES),
expand("bam/{s}.sorted.bam.bai", sample=SAMPLES),
"counts/all_samples_counts.txt",
"qc/multiqc_report.html"
# ── Step 1: Quality control ──────────────────
rule fastqc:
input: "data/{sample}.fastq.gz"
output:
html = "qc/{sample}_fastqc.html",
zip = "qc/{sample}_fastqc.zip"
threads: 4
log: "logs/fastqc/{sample}.log"
shell:
"fastqc {input} -t {threads} -o qc/ &> {log}"
# ── Step 2: Adapter trimming ─────────────────
rule trim:
input: "data/{sample}.fastq.gz"
output: "trimmed/{sample}.trimmed.fastq.gz"
threads: 4
log: "logs/trim/{sample}.log"
params:
adapters = config.get("adapters", "TruSeq3-SE.fa"),
minlen = config.get("min_length", 36)
shell:
"""
trimmomatic SE \
-threads {threads} \
{input} {output} \
ILLUMINACLIP:{params.adapters}:2:30:10 \
MINLEN:{params.minlen} \
&> {log}
"""
# ── Step 3: Alignment ────────────────────────
rule star_align:
input:
reads = "trimmed/{sample}.trimmed.fastq.gz",
index = REF
output:
bam = temp("star/{sample}/Aligned.sortedByCoord.out.bam")
threads: 8
log: "logs/star/{sample}.log"
params:
outdir = "star/{sample}/"
shell:
"""
STAR \
--runThreadN {threads} \
--genomeDir {input.index} \
--readFilesIn {input.reads} \
--readFilesCommand zcat \
--outSAMtype BAM SortedByCoordinate \
--outFileNamePrefix {params.outdir} \
&> {log}
"""
# ── Step 4: Sort and index ───────────────────
rule sort_index:
input: "star/{sample}/Aligned.sortedByCoord.out.bam"
output:
bam = "bam/{sample}.sorted.bam",
bai = "bam/{sample}.sorted.bam.bai"
threads: 4
shell:
"""
samtools sort -@ {threads} {input} -o {output.bam}
samtools index {output.bam}
"""
# ── Step 5: Feature counting ─────────────────
rule featurecounts:
input:
bams = expand("bam/{s}.sorted.bam", s=SAMPLES),
gtf = GTF
output: "counts/all_samples_counts.txt"
threads: 4
log: "logs/featurecounts.log"
shell:
"""
featureCounts \
-T {threads} \
-a {input.gtf} \
-o {output} \
{input.bams} \
&> {log}
"""
# ── Step 6: MultiQC summary ──────────────────
rule multiqc:
input:
expand("qc/{s}_fastqc.html", s=SAMPLES),
expand("logs/star/{s}.log", s=SAMPLES),
"logs/featurecounts.log"
output: "qc/multiqc_report.html"
shell: "multiqc qc/ logs/ -o qc/"
The config file that drives it:
# config.yaml
samples:
- ctrl_01
- ctrl_02
- ctrl_03
- treat_01
- treat_02
- treat_03
genome: "/scratch/references/hg38/star_index"
gtf: "/scratch/references/hg38/gencode.v44.gtf"
adapters: "TruSeq3-SE.fa"
min_length: 36
Running Snakemake
# Install
conda install -c bioconda -c conda-forge snakemake
# Dry run first — see what WOULD be executed without running anything
snakemake --dry-run
# Run on your laptop (8 cores)
snakemake --cores 8
# Run on HPC via SLURM (Snakemake submits each rule as a SLURM job)
snakemake --profile slurm --jobs 50
# Visualise the DAG (requires graphviz)
snakemake --dag | dot -Tpng > dag.png
# Force rerun everything from scratch
snakemake --cores 8 --forceall
# Rerun only rules that depend on a changed file
snakemake --cores 8 --rerun-incomplete
Running on HPC with a SLURM Profile
Create a SLURM profile directory:
mkdir -p ~/.config/snakemake/slurm/
cat > ~/.config/snakemake/slurm/config.yaml << 'EOF'
executor: slurm
jobs: 100
latency-wait: 60
default-resources:
slurm_partition: normal
mem_mb: 8192
runtime: 60
cpus_per_task: 4
EOF
Now snakemake --profile slurm --jobs 50 automatically submits each rule as its own SLURM job, with the threads: value used as --cpus-per-task.
Part 3: Nextflow — Cloud-Native and Scalable
Nextflow uses a different philosophy: instead of rules, it defines processes that communicate through channels. It’s more verbose than Snakemake but scales better to very large pipelines and is the preferred choice in industry and large consortia (nf-core, Seqera).
Core Concept: Processes and Channels
// The most basic Nextflow process
process FASTQC {
input: path reads
output: path "*.html"
script:
"""
fastqc $reads
"""
}
A channel is a stream of data that flows into processes:
workflow {
// Create a channel from all FASTQ files
reads_ch = Channel.fromPath("data/*.fastq.gz")
// Pass the channel to the process
FASTQC(reads_ch)
}
A Complete RNA-seq Nextflow Pipeline
// main.nf — Nextflow DSL2 RNA-seq pipeline
// ─────────────────────────────────────────
nextflow.enable.dsl=2
// ── Parameters ───────────────────────────
params.reads = "data/*_{R1,R2}.fastq.gz"
params.genome = "/scratch/references/hg38/star_index"
params.gtf = "/scratch/references/hg38/gencode.v44.gtf"
params.outdir = "results"
// ── Process: FastQC ───────────────────────
process FASTQC {
tag "$sample_id"
publishDir "${params.outdir}/qc", mode: 'copy'
input:
tuple val(sample_id), path(reads)
output:
tuple val(sample_id), path("*.html"), path("*.zip")
script:
"""
fastqc $reads -t $task.cpus
"""
}
// ── Process: Trim ─────────────────────────
process TRIM {
tag "$sample_id"
publishDir "${params.outdir}/trimmed", mode: 'copy'
input:
tuple val(sample_id), path(reads)
output:
tuple val(sample_id), path("${sample_id}.trimmed_{R1,R2}.fastq.gz")
script:
"""
trimmomatic PE \
-threads $task.cpus \
${reads[0]} ${reads[1]} \
${sample_id}.trimmed_R1.fastq.gz /dev/null \
${sample_id}.trimmed_R2.fastq.gz /dev/null \
ILLUMINACLIP:TruSeq3-PE.fa:2:30:10 MINLEN:36
"""
}
// ── Process: STAR alignment ───────────────
process STAR {
tag "$sample_id"
publishDir "${params.outdir}/bam", mode: 'copy'
cpus 8
memory '32 GB'
input:
tuple val(sample_id), path(reads)
output:
tuple val(sample_id), path("${sample_id}.sorted.bam"),
path("${sample_id}.sorted.bam.bai")
script:
"""
STAR \
--runThreadN $task.cpus \
--genomeDir ${params.genome} \
--readFilesIn ${reads[0]} ${reads[1]} \
--readFilesCommand zcat \
--outSAMtype BAM SortedByCoordinate \
--outFileNamePrefix ${sample_id}_
samtools sort -@ $task.cpus ${sample_id}_Aligned.sortedByCoord.out.bam \
-o ${sample_id}.sorted.bam
samtools index ${sample_id}.sorted.bam
"""
}
// ── Process: featureCounts ────────────────
process FEATURECOUNTS {
publishDir "${params.outdir}/counts", mode: 'copy'
input:
path bams
output:
path "all_counts.txt"
script:
"""
featureCounts \
-T $task.cpus \
-a ${params.gtf} \
-o all_counts.txt \
$bams
"""
}
// ── Workflow definition ───────────────────
workflow {
// Create channel of paired-end reads
reads_ch = Channel
.fromFilePairs(params.reads)
.ifEmpty { error "No reads found: ${params.reads}" }
// Run pipeline
fastqc_out = FASTQC(reads_ch)
trimmed = TRIM(reads_ch)
aligned = STAR(trimmed)
// Collect all BAMs for featureCounts
bam_files = aligned.map { id, bam, bai -> bam }.collect()
FEATURECOUNTS(bam_files)
}
Running Nextflow
# Install
conda install -c bioconda nextflow
# Test run (no data needed)
nextflow run hello
# Run your pipeline locally
nextflow run main.nf
# Run on HPC with SLURM
nextflow run main.nf -profile slurm
# Resume from last successful step after a failure
nextflow run main.nf -resume
# Run with custom parameters
nextflow run main.nf \
--reads "data/*_{R1,R2}.fastq.gz" \
--genome /scratch/refs/hg38 \
--outdir my_results
Create a SLURM execution profile in nextflow.config:
// nextflow.config
profiles {
slurm {
process.executor = 'slurm'
process.queue = 'normal'
process.clusterOptions = '--account=myproject'
process {
withName: STAR {
cpus = 8
memory = '32 GB'
time = '4h'
}
withName: FASTQC {
cpus = 4
memory = '8 GB'
time = '1h'
}
}
}
}
Part 4: Snakemake vs Nextflow — Choosing Your Tool
| Feature | Snakemake | Nextflow |
|---|---|---|
| Language | Python | Groovy / DSL2 |
| Learning curve | Lower (if you know Python) | Higher |
| Syntax style | Rule-based (input → output) | Process + channel streams |
| HPC support | Excellent (profiles) | Excellent (executors) |
| Cloud support | Good | Excellent (AWS, GCP, Azure) |
| Resuming failed jobs | --rerun-incomplete | -resume (automatic) |
| Community pipelines | Many | nf-core (industry standard) |
| Best for | Academic labs, Python users | Large consortia, industry, cloud |
| Debugging | Generally easier | Steeper learning curve |
My recommendation:
- New to workflows? → Start with Snakemake. The Python syntax is readable, the documentation is excellent, and it will handle everything in a typical academic bioinformatics project.
- Large collaborative project / cloud / sharing with the community? → Nextflow + nf-core. The nf-core library has production-ready pipelines for RNA-seq, metagenomics, variant calling, and 100+ other workflows that you can run with a single command.
# Use a production nf-core pipeline (no writing code required)
nextflow run nf-core/rnaseq \
--input samplesheet.csv \
--genome GRCh38 \
--outdir results \
-profile singularity,slurm
Part 5: GitHub Repository Structure
A reproducible pipeline is only useful if someone else can run it. Here’s the repository structure that makes that possible:
my-rnaseq-pipeline/
├── README.md ← setup + usage instructions
├── config.yaml ← parameters (samples, paths, tool settings)
├── environment.yml ← exact conda environment (from Day 2)
├── Snakefile ← pipeline definition
│
├── data/ ← symlinks to raw data (never commit raw data)
│ └── .gitkeep
│
├── resources/
│ ├── TruSeq3-SE.fa ← adapter sequences
│ └── genome_paths.yaml ← cluster-specific reference paths
│
├── logs/ ← SLURM job logs (gitignored)
│ └── .gitignore
│
├── results/ ← pipeline outputs (gitignored)
│ └── .gitignore
│
└── scripts/
├── setup_env.sh ← create conda env + download references
└── run_pipeline.sh ← one-command launch script
The run_pipeline.sh wrapper makes it trivial to reproduce:
#!/bin/bash
# run_pipeline.sh — one command to reproduce the analysis
set -euo pipefail
conda activate rnaseq-pipeline
snakemake --profile slurm --jobs 100 --rerun-incomplete
Your README should answer four questions in under 60 seconds:
## Quick Start
**1. Clone and set up environment**
git clone https://github.com/you/my-rnaseq-pipeline
conda env create -f environment.yml
conda activate rnaseq-pipeline
**2. Edit config.yaml**
- Add your sample names
- Update genome/GTF paths for your cluster
**3. Run**
bash scripts/run_pipeline.sh
**Expected output:** results/counts/all_samples_counts.txt
**Runtime:** ~2 hours on 6 samples with SLURM
Part 6: Reproducibility Checklist
Before you share your pipeline or submit a paper:
□ All software versions pinned in environment.yml
□ All parameters in config.yaml (not hardcoded in scripts)
□ All paths are relative or configurable (no /home/yourname/ hardcoded)
□ README answers: install, configure, run, expected output
□ Snakefile/main.nf is committed to Git
□ At least one test dataset included or linked
□ Results directory is in .gitignore
□ Each rule/process has a log: directive
□ Pipeline tested with --dry-run before full submission
□ seff output recorded for at least one full run (resource documentation)
Part 7: The Full Series Recap
You’ve now covered the complete arc from laptop to reproducible HPC pipeline:
| Day | Topic | Key Skill |
|---|---|---|
| 1 | Laptop vs HPC | Understanding the landscape |
| 2 | Software installation | conda, modules, environment.yml |
| 3 | Running jobs | SLURM, sbatch, %j, interactive nodes |
| 4 | Scaling | Job arrays, $SLURM_ARRAY_TASK_ID |
| 5 | Pipelines | Snakemake, Nextflow, reproducibility |
| 6 | Data transfer | scp, rsync, Globus Connect |
The through-line across all six posts: your bioinformatics tools don’t change. fastqc, bowtie2, samtools, kraken2 — they all work the same. What changes is the infrastructure around them: how you install them, how you run them, how you scale them, and how you make them reproducible.
Common Mistakes When Starting With Workflows
Putting the whole pipeline in rule all
# ❌ This defeats the purpose
rule all:
shell: "bash my_manual_script.sh"
# ✅ Define inputs and outputs properly so Snakemake can track them
rule all:
input: expand("results/{s}.bam", s=SAMPLES)
Hardcoding paths
# ❌ Only works on your machine
input: "/home/yourname/project/data/{sample}.fastq.gz"
# ✅ Use config or relative paths
input: config["data_dir"] + "/{sample}.fastq.gz"
# or simply:
input: "data/{sample}.fastq.gz"
Not using log: directives
# ❌ Errors go to the terminal and get lost
rule fastqc:
shell: "fastqc {input}"
# ✅ Always capture logs
rule fastqc:
log: "logs/fastqc/{sample}.log"
shell: "fastqc {input} &> {log}"
Starting with Nextflow before understanding the concept
Start with Snakemake. Get one pipeline working end-to-end. Understand the DAG concept. Then explore Nextflow when your project outgrows what Snakemake handles comfortably.
Try It Yourself
# 1. Install Snakemake
conda activate bioenv
conda install -c bioconda -c conda-forge snakemake -y
# 2. Create a minimal test Snakefile
mkdir -p snakemake_test/{data,qc,logs}
cd snakemake_test/
# Create a dummy FASTQ
echo -e "@r1\nACGT\n+\nIIII" > data/sample1.fastq.gz
cat > Snakefile << 'EOF'
SAMPLES = ["sample1"]
rule all:
input: expand("qc/{s}_fastqc.html", s=SAMPLES)
rule fastqc:
input: "data/{sample}.fastq.gz"
output: "qc/{sample}_fastqc.html"
log: "logs/{sample}_fastqc.log"
shell: "fastqc {input} -o qc/ &> {log}"
EOF
# 3. Dry run
snakemake --dry-run
# 4. Run
snakemake --cores 2
# 5. Nothing changed — it won't rerun
snakemake --cores 2
# Nothing to be done.
Summary
- Shell scripts don’t track dependencies — workflow managers do
- Snakemake uses Python-style rules (input → output → shell). Best for Python users and academic projects
- Nextflow uses DSL2 processes + channels. Best for large-scale, cloud, and community pipelines (nf-core)
- Both integrate with SLURM: submit each step as its own HPC job automatically
- A good GitHub repository structure + README makes your pipeline reproducible by anyone
- The
-resume(Nextflow) and--rerun-incomplete(Snakemake) flags mean a failed job never means rerunning everything
Up Next
Day 6: Moving Your Data — scp, rsync, and Globus Connect
Your pipeline is reproducible. Now: how do you get 300 GB of raw reads onto the cluster in the first place? We cover every data transfer method you’ll use in practice — from quick one-liners to TB-scale transfers that run unattended.