Repositories
Research code, bioinformatics pipelines, and reproducible workflows.
🧬 Research Repositories
This page provides an overview of code repositories, pipelines, and computational resources I maintain or actively develop. My repositories emphasize reproducibility, scalability, and clarity, with workflows designed for HPC environments and real-world biological datasets.
Most projects are hosted on GitHub:
🔗 GitHub: https://github.com/jojyjohn28
🔗 Blog for more details : https://jojyjohn28.github.io/blog/
🧪 Amplicon Analysis
🔗 https://github.com/jojyjohn28/AmpliconWeek_2025
A structured, day-by-day workflow for amplicon sequencing analysis, developed as both a training resource and a reproducible analysis guide.
Focus areas:
- QIIME2-based amplicon processing
- Diversity analysis and visualization
- Ecological interpretation of ASVs
- Best practices for reproducible workflows

🧬 Genome-Resolved Metagenomics
🔗 https://github.com/jojyjohn28/semibin2-soil-mag-workflow
A Snakemake-based workflow for SemiBin2-assisted MAG recovery, optimized for soil and complex environmental metagenomes.
Focus areas:
- Co-assembly and binning strategies
- Semi-supervised MAG recovery
- Genome quality assessment
- Scalable execution on HPC systems
🧬 Metagenome Analysis Series
🔗 https://github.com/jojyjohn28/metagenome-analysis-series
A comprehensive, hands-on tutorial series for analyzing metagenomic data from raw reads to biological insights. This series covers the complete workflow used in modern metagenomics research, from quality control to advanced downstream analyses.
Focus areas: 📖 Day 1: QC & Taxonomic Profiling
📖 Day 2: Genome Assembly
📖 Day 3: Genome Binning
📖 Day 4: Dereplication & Taxonomy
📖 Day 5: Genome Annotation
📖 Day 6: Specialized Functions
📖 Day 7: Comparative Genomics
📖 Day 8: Workflow Platforms
📖 Day 9: Visualization
📖 Day 10: Multi-Omics Integration
Files, toy data and codes are available 🔗 Course Materials GitHub
🔗 10-Day Series GitHub 🔗 10-Day Series blog

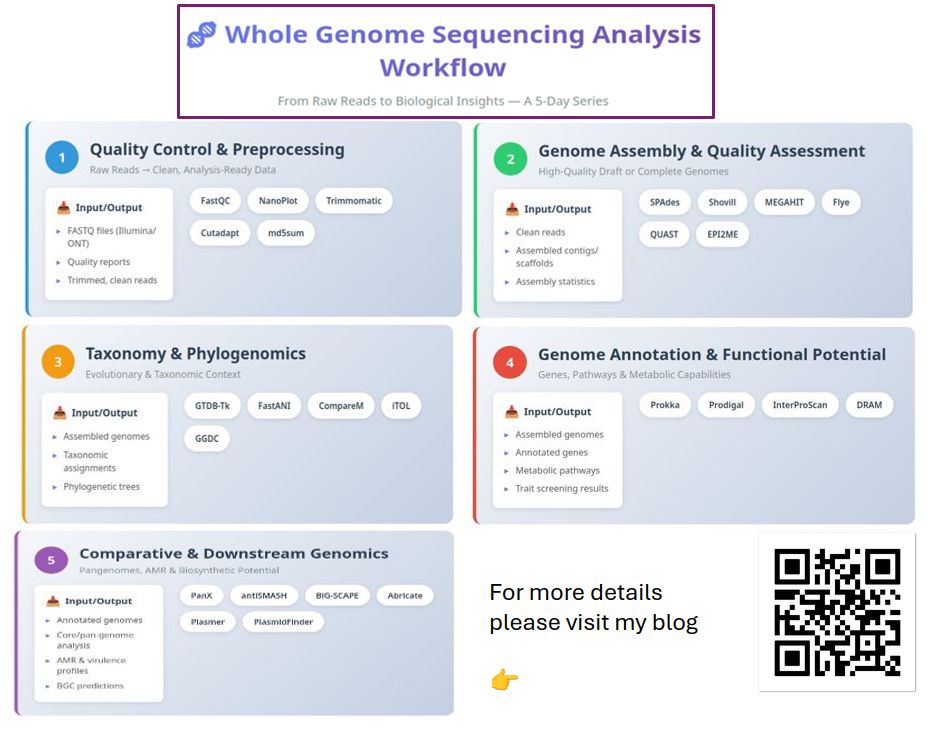
🧬 Whole Genome Sequencing Analysis — From Raw Reads to Biological Insight
https://github.com/jojyjohn28/whole-genome-sequencing-analysis
This repository documents a step-by-step, reproducible whole-genome sequencing (WGS) analysis workflow, developed and applied to bacterial genomes generated from both Illumina (short-read) and Oxford Nanopore (long-read) sequencing.
Focus areas: Day 1 — Raw Reads to Clean, Analysis-Ready Data Day 2 — Genome Assembly & Assembly Quality Assessment Day 3 — Taxonomy, Phylogeny & Genome Similarity Day 4 — Genome Annotation & Functional Potential Day 5 — Comparative & Downstream Genomic Analyses
🧬 ML-heart-to-ecosystem 🫀🌱
https://github.com/jojyjohn28/ML-heart-to-ecosystem
A comprehensive machine learning journey from clinical heart disease prediction to ecosystem multifunctionality analysis. Validating Random Forest and ensemble methods on medical data before applying them to predict soil ecosystem functions from microbial communities.
Focus areas:
-
Heart Disease Prediction (Clinical ML) - Complete ✅ ●303 patients, 14 clinical features ● 5 algorithms compared: Logistic Regression, Random Forest, XGBoost, SVM, KNN ● Best Result: 88.52% accuracy, 91.18% recall (Logistic Regression)
-
Soil Ecosystem Multifunctionality (Ecological ML) - In Progress 🚧 ● Several soil samples across different cover crops ● Predicting ecosystem functions from microbial communities ● Applying validated Random Forest methods
🧬 Functional Redundancy and Metabolic Flexibility of Estuarine Microbiomes
https://github.com/jojyjohn28/FRed-estuarine-microbiome
This repository contains data, code, and figures supporting the manuscript:”Functional redundancy and metabolic flexibility of estuarine microbiomes” Read more at : https://academic.oup.com/ismecommun/article/6/1/ycag021/8454622

🌊 Size_Fractionated_Microbiome_Analysis**
🔗 https://github.com/jojyjohn28/Size_Fractionated_Microbiome_Analysis
A comprehensive analysis framework for free-living and particle-associated microbial communities, integrating metagenomics and metatranscriptomics.
Focus areas:
- Read-based taxonomic and functional profiling
- DNA–RNA comparisons (total vs active communities)
- Resource partitioning and substrate uptake
- Functional redundancy (FRed) analysis
- Co-occurrence and ecological interpretation
ongoing work
Across all repositories, I emphasize:
- Reproducibility over one-off scripts
- Clear documentation and modular design
- Compatibility with HPC environments (SLURM)
- Teaching workflows that users can reproduce and troubleshoot independently
These repositories are used in research projects, student mentoring, and collaborative studies.
📌 Notes
- Repositories may changes as the work progress.!
- Please visit : https://github.com/jojyjohn28 for updates